Home > Press > Scientists Map Key Moment in Assembly of DNA-Splitting Molecular Machine: Crucial steps and surprising structures revealed in the genesis of the enzyme that divides the DNA double helix during cell replication
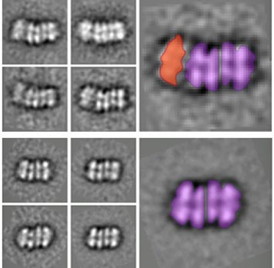 |
| Averaged electron microscope images of two intermediate helicase structures. The top shows an ORC binding the two ring structures together, and the bottom shows the completed double-ring structure with the ORC detached. |
Abstract:
The proteins that drive DNA replication-the force behind cellular growth and reproduction-are some of the most complex machines on Earth. The multistep replication process involves hundreds of atomic-scale moving parts that rapidly interact and transform. Mapping that dense molecular machinery is one of the most promising and challenging frontiers in medicine and biology.
Scientists Map Key Moment in Assembly of DNA-Splitting Molecular Machine: Crucial steps and surprising structures revealed in the genesis of the enzyme that divides the DNA double helix during cell replication
Upton, NY | Posted on October 15th, 2014Now, scientists have pinpointed crucial steps in the beginning of the replication process, including surprising structural details about the enzyme that "unzips" and splits the DNA double helix so the two halves can serve as templates for DNA duplication.
The research combined electron microscopy, perfectly distilled proteins, and a method of chemical freezing to isolate specific moments at the start of replication. The study-authored by scientists from the U.S. Department of Energy's Brookhaven National Laboratory, Stony Brook University, Cold Spring Harbor Laboratory, and Imperial College, London-published on Oct. 15, 2014, in the journal Genes and Development.
"The genesis of the DNA-unwinding machinery is wonderfully complex and surprising," said study coauthor Huilin Li, a biologist at Brookhaven Lab and Stony Brook University. "Seeing this helicase enzyme prepare to surround and unwind the DNA at the molecular level helps us understand the most fundamental process of life and how that process might go wrong. Errors in copying DNA are found in certain cancers, and this work could one day help develop new treatment methods that stall or break dangerous runaway machinery."
The research picks up where two previous studies by Li and colleagues left off. They first determined the structure of the "Origin Recognition Complex" (ORC), a protein that identifies and attaches to specific DNA sites to initiate the entire replication process. The second study revealed how the ORC recruits, cracks open, and installs a crucial ring-shaped protein structure (Mcm2-7) that lies at the core of the helicase enzyme.
But DNA replication is a bi-directional process with two helicases moving in opposite directions. The key question, then, was how does a second helicase core get recruited and loaded onto the DNA in the opposite orientation of the first?
"To our surprise, we found an intermediate structure with one ORC binding two rings," said Brookhaven Lab biologist and lead author Jingchuan Sun. "This discovery suggests that a single ORC, rather than the commonly believed two-ORC system, loads both helicase rings."
One step further along, the researchers also determined the molecular architecture of the final double-ring structure left behind after the ORC leaves the system, offering a number of key biological insights.
"We now have clues to how that double-ring structure stably lingers until the cell enters the DNA-synthesis phase much later on in replication," said study coauthor Christian Speck of Imperial College, London. "This study revealed key regulatory principles that explain how the helicase activity is initially suppressed and then becomes reactivated to begin its work splitting the DNA."
Precision methods, close collaboration
Examining these fleeting molecular structures required mastery of biology, chemistry, and electron microscopy techniques.
"This three-way collaboration took advantage of each lab's long standing collaboration and expertise," said study coauthor Bruce Stillman of Cold Spring Harbor. "Imperial College and Cold Spring Harbor handled the challenging material preparation and functional characterization, while Brookhaven and Stony Brook led the sophisticated molecular imaging and three-dimensional image reconstruction."
The researchers used proteins from baker's yeast-a model organism for the more complex systems found in animals. The scientists isolated the protein mechanisms involved in replication and removed structures that might otherwise complicate the images.
Once the isolated proteins were mixed with DNA, the scientists injected chemicals to "freeze" the binding and recruitment process at intervals of 2, 7, and 30 minutes.
They then used an electron microscope at Brookhaven to pin down the exact structures at each targeted moment in a kind of molecular time-lapse. Rather than the light used in a traditional microscope, this technique uses focused beams of electrons to illuminate a sample and form images with atomic resolution. The instrument produces a large number of two-dimensional electron beam images, which a computer then reconstructs into three-dimensional structure.
"This technique is ideal because we're imaging relatively massive proteins here," Li said. "A typical protein contains three hundred amino acids, but these DNA replication mechanisms consist of tens of thousands of amino acids. The entire structure is about 20-nanometers across, compared to 4 nanometers for an average protein."
Unraveling the DNA processes at the most fundamental level, the focus of this team's work, could have far-reaching implications.
"The structural knowledge may help others engineer small molecules that inhibit DNA replication at specific moments, leading to new disease prevention or treatment techniques," Li said.
Additional collaborators on this research include Alejandra Fernandez, Alberto Riera, and Silvia Tognetti of the MRC Clinical Science Centre of Imperial College, London; and Zuanning Yuan of Stony Brook University.
The research was funded by the National Institutes of Health (GM45436, GM74985) and the United Kingdom Medical Research Council.
Brookhaven National Laboratory is supported by the Office of Science of the U.S. Department of Energy. The Office of Science is the single largest supporter of basic research in the physical sciences in the United States, and is working to address some of the most pressing challenges of our time. For more information, please visit science.energy.gov.
####
About Brookhaven National Laboratory
One of ten national laboratories overseen and primarily funded by the Office of Science of the U.S. Department of Energy (DOE), Brookhaven National Laboratory conducts research in the physical, biomedical, and environmental sciences, as well as in energy technologies and national security. Brookhaven Lab also builds and operates major scientific facilities available to university, industry and government researchers. Brookhaven is operated and managed for DOE's Office of Science by Brookhaven Science Associates, a limited-liability company founded by the Research Foundation for the State University of New York on behalf of Stony Brook University, the largest academic user of Laboratory facilities, and Battelle, a nonprofit applied science and technology organization.
For more information, please click here
Contacts:
Justin Eure
(631) 344-2347
or
Peter Genzer
(631) 344-3174
Copyright © Brookhaven National Laboratory
If you have a comment, please Contact us.Issuers of news releases, not 7th Wave, Inc. or Nanotechnology Now, are solely responsible for the accuracy of the content.
| Related News Press |
News and information
![]() Quantum computer improves AI predictions April 17th, 2026
Quantum computer improves AI predictions April 17th, 2026
![]() Flexible sensor gains sensitivity under pressure April 17th, 2026
Flexible sensor gains sensitivity under pressure April 17th, 2026
![]() A reusable chip for particulate matter sensing April 17th, 2026
A reusable chip for particulate matter sensing April 17th, 2026
![]() Detecting vibrational quantum beating in the predissociation dynamics of SF6 using time-resolved photoelectron spectroscopy April 17th, 2026
Detecting vibrational quantum beating in the predissociation dynamics of SF6 using time-resolved photoelectron spectroscopy April 17th, 2026
Laboratories
![]() Researchers develop molecular qubits that communicate at telecom frequencies October 3rd, 2025
Researchers develop molecular qubits that communicate at telecom frequencies October 3rd, 2025
![]() A battery�s hopping ions remember where they�ve been: Seen in atomic detail, the seemingly smooth flow of ions through a battery�s electrolyte is surprisingly complicated February 16th, 2024
A battery�s hopping ions remember where they�ve been: Seen in atomic detail, the seemingly smooth flow of ions through a battery�s electrolyte is surprisingly complicated February 16th, 2024
Govt.-Legislation/Regulation/Funding/Policy
![]() Quantum computer improves AI predictions April 17th, 2026
Quantum computer improves AI predictions April 17th, 2026
![]() Metasurfaces smooth light to boost magnetic sensing precision January 30th, 2026
Metasurfaces smooth light to boost magnetic sensing precision January 30th, 2026
![]() New imaging approach transforms study of bacterial biofilms August 8th, 2025
New imaging approach transforms study of bacterial biofilms August 8th, 2025
Nanomedicine
![]() A fundamentally new therapeutic approach to cystic fibrosis: Nanobody repairs cellular defect April 17th, 2026
A fundamentally new therapeutic approach to cystic fibrosis: Nanobody repairs cellular defect April 17th, 2026
![]() New molecular technology targets tumors and simultaneously silences two �undruggable� cancer genes August 8th, 2025
New molecular technology targets tumors and simultaneously silences two �undruggable� cancer genes August 8th, 2025
![]() New imaging approach transforms study of bacterial biofilms August 8th, 2025
New imaging approach transforms study of bacterial biofilms August 8th, 2025
![]() Electrifying results shed light on graphene foam as a potential material for lab grown cartilage June 6th, 2025
Electrifying results shed light on graphene foam as a potential material for lab grown cartilage June 6th, 2025
Discoveries
![]() Quantum computer improves AI predictions April 17th, 2026
Quantum computer improves AI predictions April 17th, 2026
![]() Flexible sensor gains sensitivity under pressure April 17th, 2026
Flexible sensor gains sensitivity under pressure April 17th, 2026
![]() A reusable chip for particulate matter sensing April 17th, 2026
A reusable chip for particulate matter sensing April 17th, 2026
![]() Detecting vibrational quantum beating in the predissociation dynamics of SF6 using time-resolved photoelectron spectroscopy April 17th, 2026
Detecting vibrational quantum beating in the predissociation dynamics of SF6 using time-resolved photoelectron spectroscopy April 17th, 2026
Announcements
![]() A fundamentally new therapeutic approach to cystic fibrosis: Nanobody repairs cellular defect April 17th, 2026
A fundamentally new therapeutic approach to cystic fibrosis: Nanobody repairs cellular defect April 17th, 2026
![]() UC Irvine physicists discover method to reverse �quantum scrambling� : The work addresses the problem of information loss in quantum computing system April 17th, 2026
UC Irvine physicists discover method to reverse �quantum scrambling� : The work addresses the problem of information loss in quantum computing system April 17th, 2026
Interviews/Book Reviews/Essays/Reports/Podcasts/Journals/White papers/Posters
![]() A fundamentally new therapeutic approach to cystic fibrosis: Nanobody repairs cellular defect April 17th, 2026
A fundamentally new therapeutic approach to cystic fibrosis: Nanobody repairs cellular defect April 17th, 2026
![]() UC Irvine physicists discover method to reverse �quantum scrambling� : The work addresses the problem of information loss in quantum computing system April 17th, 2026
UC Irvine physicists discover method to reverse �quantum scrambling� : The work addresses the problem of information loss in quantum computing system April 17th, 2026
Nanobiotechnology
![]() A fundamentally new therapeutic approach to cystic fibrosis: Nanobody repairs cellular defect April 17th, 2026
A fundamentally new therapeutic approach to cystic fibrosis: Nanobody repairs cellular defect April 17th, 2026
![]() New molecular technology targets tumors and simultaneously silences two �undruggable� cancer genes August 8th, 2025
New molecular technology targets tumors and simultaneously silences two �undruggable� cancer genes August 8th, 2025
![]() New imaging approach transforms study of bacterial biofilms August 8th, 2025
New imaging approach transforms study of bacterial biofilms August 8th, 2025
![]() Electrifying results shed light on graphene foam as a potential material for lab grown cartilage June 6th, 2025
Electrifying results shed light on graphene foam as a potential material for lab grown cartilage June 6th, 2025
Research partnerships
![]() Lab to industry: InSe wafer-scale breakthrough for future electronics August 8th, 2025
Lab to industry: InSe wafer-scale breakthrough for future electronics August 8th, 2025
![]() HKU physicists uncover hidden order in the quantum world through deconfined quantum critical points April 25th, 2025
HKU physicists uncover hidden order in the quantum world through deconfined quantum critical points April 25th, 2025
|
|
||
|
|
||
| The latest news from around the world, FREE | ||
|
|
||
|
|
||
| Premium Products | ||
|
|
||
|
Only the news you want to read!
Learn More |
||
|
|
||
|
Full-service, expert consulting
Learn More |
||
|
|
||